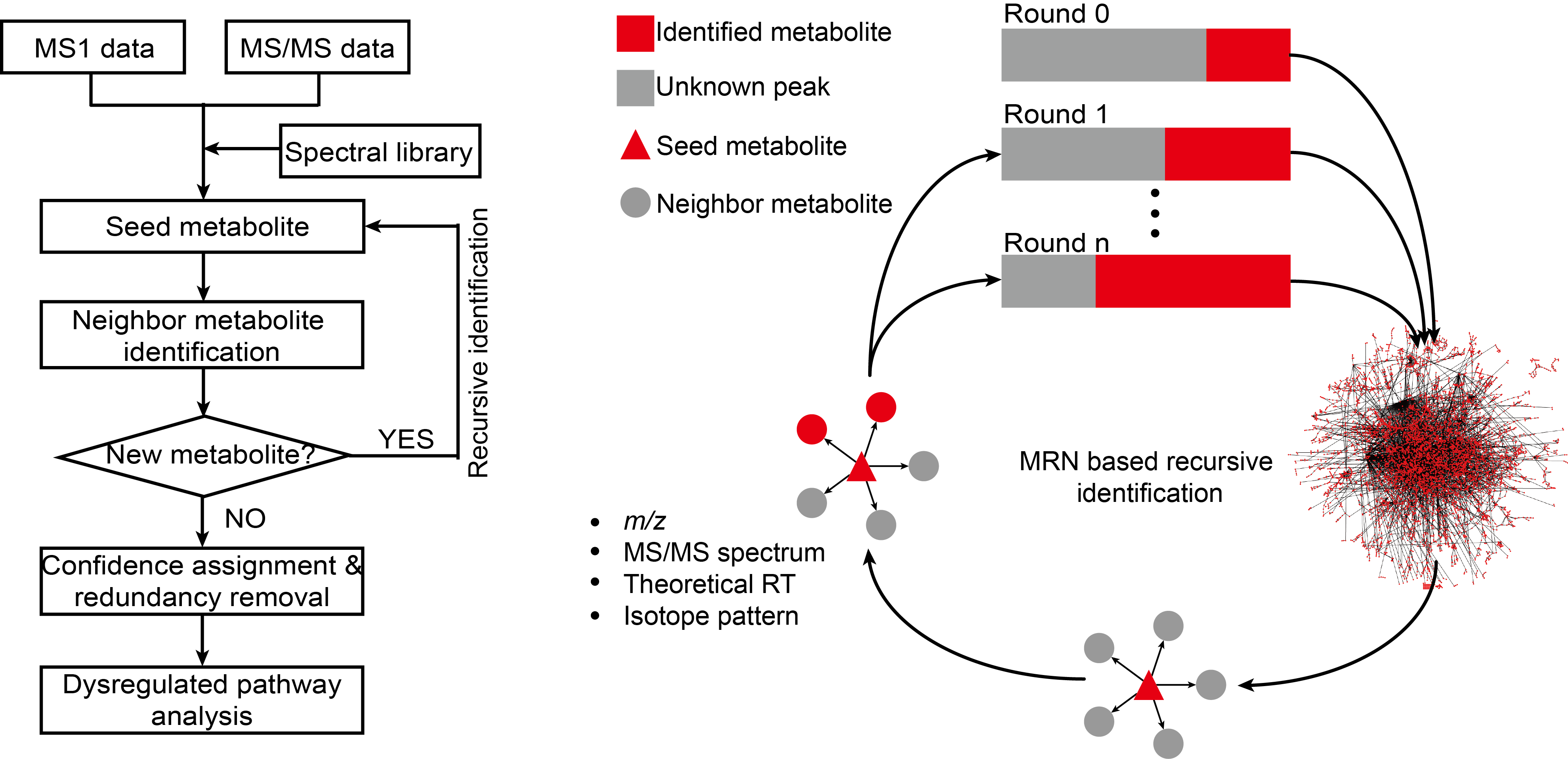
估计阅读时长: 6 分钟访问在线服务: http://metdna.zhulab.cn/ Metabolite identification is the long-standing challenge for liquid chromatography-mass spectrometry (LC-MS)-based untargeted metabolomics. Here, […]
Recent Posts
Archives
- February 2026 (9)
- January 2026 (2)
- December 2025 (10)
- November 2025 (2)
- October 2025 (1)
- August 2025 (3)
- July 2025 (2)
- June 2025 (6)
- May 2025 (3)
- November 2023 (1)
- June 2023 (2)
- May 2023 (2)
- April 2023 (2)
- March 2023 (2)
- February 2023 (1)
- August 2022 (2)
- July 2022 (2)
- June 2022 (5)
- May 2022 (5)
- April 2022 (4)
- March 2022 (3)
- January 2022 (2)
- December 2021 (2)
- November 2021 (2)
- October 2021 (6)
- September 2021 (8)
- August 2021 (8)
- July 2021 (6)
- June 2021 (20)
- May 2021 (10)
博客文章
| S | M | T | W | T | F | S |
|---|---|---|---|---|---|---|
| 1 | 2 | |||||
| 3 | 4 | 5 | 6 | 7 | 8 | 9 |
| 10 | 11 | 12 | 13 | 14 | 15 | 16 |
| 17 | 18 | 19 | 20 | 21 | 22 | 23 |
| 24 | 25 | 26 | 27 | 28 | 29 | 30 |
| 31 | ||||||
Tags
algorithm (33)
bilibili (3)
binary tree (4)
clustering (21)
Darwinism (4)
data visualization (23)
dotnet-core (25)
ec number (4)
embedding (5)
GCModeller (20)
gdi+ (23)
gem (7)
ggplot (14)
graph (14)
heatmap (5)
http (4)
image processing (7)
kegg (8)
kmeans (3)
language (7)
linq (3)
linux (8)
machine learning (4)
mass spectrometry (12)
math (19)
metagenomics (6)
motif (4)
MSI (4)
mzkit (19)
network (8)
pathway (4)
pipeline (4)
query (5)
R# (44)
rsharp (23)
scripting (14)
single-cell (6)
sql (3)
symbolic computation (3)
text processing (4)
typescript (3)
ubuntu (4)
uniprot (3)
vb (19)
VisualBasic (50)


[…] 我们在基于前面所论述的《通过diamond软件进行blastp搜索》对大规模的基因组数据进行了代谢酶的EC number的注释以及按照文章《基因组功能注释(EC Number)的向量化嵌入》的方法,得到了一个比较大的基因组代谢酶TF-IDF嵌入丰度矩阵后,如果将这里所得到的嵌入结果矩阵中的基因组,基于Family层级的物种分类分组看作为单细胞转录数据中的细胞分群结果,能否基于单细胞数据分析方法来分析和可视化我的基因组功能嵌入的结果矩阵呢? […]
[…] 我们在基于前面所论述的《通过diamond软件进行blastp搜索》对大规模的基因组数据进行了代谢酶的EC number的注释以及按照文章《基因组功能注释(EC Number)的向量化嵌入》的方法,得到了一个比较大的基因组代谢酶TF-IDF嵌入丰度矩阵后,如果将这里所得到的嵌入结果矩阵中的基因组,基于Family层级的物种分类分组看作为单细胞转录数据中的细胞分群结果,能否基于单细胞数据分析方法来分析和可视化我的基因组功能嵌入的结果矩阵呢? […]
[…] 对于基于ec number来生成层级数据,我们直接使用《酶EC编号结构解析》文章末尾所展示的层级数据生成函数来实现。 […]
[…] 在前面的一篇《基因组功能注释(EC Number)的向量化嵌入》博客文章中,针对所注释得到的微生物基因组代谢信息,进行基于TF-IDF的向量化嵌入之后。为了可视化向量化嵌入的效果,通过UMAP进行降维,然后基于降维的结果进行散点图可视化。通过散点图可视化可以发现向量化的嵌入结果可以比较好的将不同物种分类来源的微生物基因组区分开来。 […]
😲啊?